
The fruit fly Drosophila melanogaster has been used to study immunity and blood cell differentiation for the past 60 years. Although, according to our recent knowledge, Drosophila lacks adaptive immunity, the functions of its immune cells (the hemocytes), such as phagocytosis, encapsulation and coagulation are similar to those of vertebrate myeloid cells. The signaling pathways, as well as the transcription and epigenetic factors that regulate blood cell differentiation are highly conserved. Similarly to vertebrate blood cells, hemocytes differentiate in spatially separated hematopoietic compartments in multiple waves. Therefore, Drosophila is an ideal model organism to study the regulation of blood cell differentiation, as well as tumor formation caused by misregulation of blood cell fate.
1. Modeling leukemia pathogenesis in Drosophila melanogaster
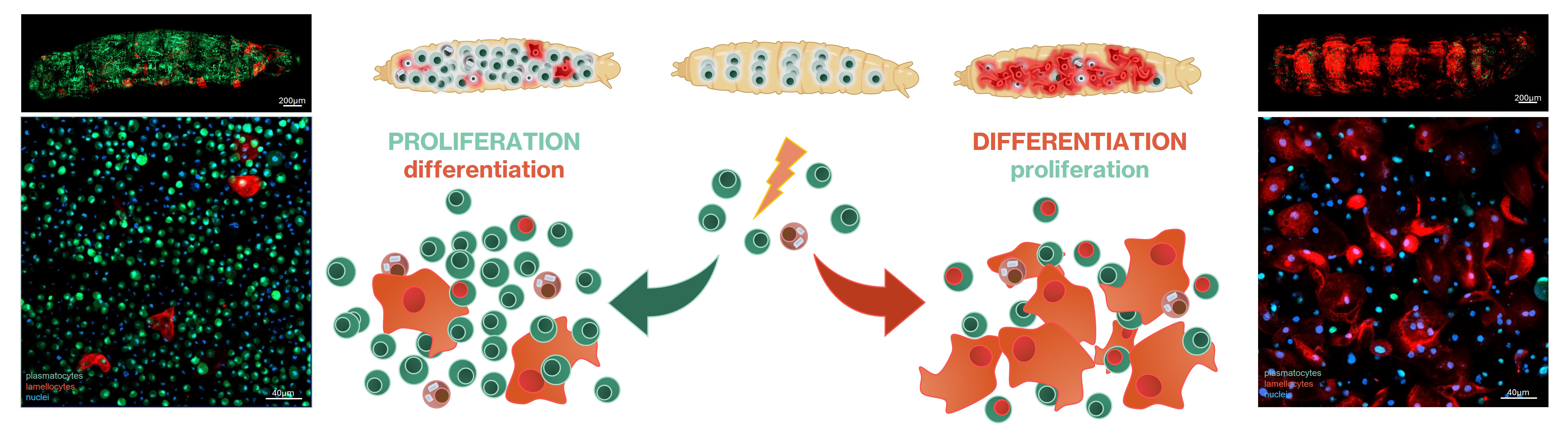
Transdifferentiation is the conversion of an already differentiated cell type into another cell type without the involvement of stem cells. Our transgenic cell lineage tracing experiments revealed that the phagocytic plasmatocytes of the larva are plastic cells that are capable of differentiating into capsule-forming lamellocytes upon immune induction.
In both mammalian and Drosophila models, blood cells exhibit uncontrolled proliferation and transdifferentiation under tumorigenic conditions. In leukemia models across diverse genetic backgrounds, the relative contributions of proliferation and transdifferentiation vary, as does the size and morphology of lamellocytes. Our objective is to identify the factors and molecular mechanisms regulating the proliferation–transdifferentiation balance and driving cellular heterogeneity in leukemia.
2. Modeling leukemia pathogenesis and developing novel therapy using human cell lines
Disruption of normal blood cell regulation can lead to malignant diseases, including leukemias and myeloproliferative neoplasms. In these disorders, blood forming cells grow uncontrollably and the mechanisms that normally regulate their function become impaired.
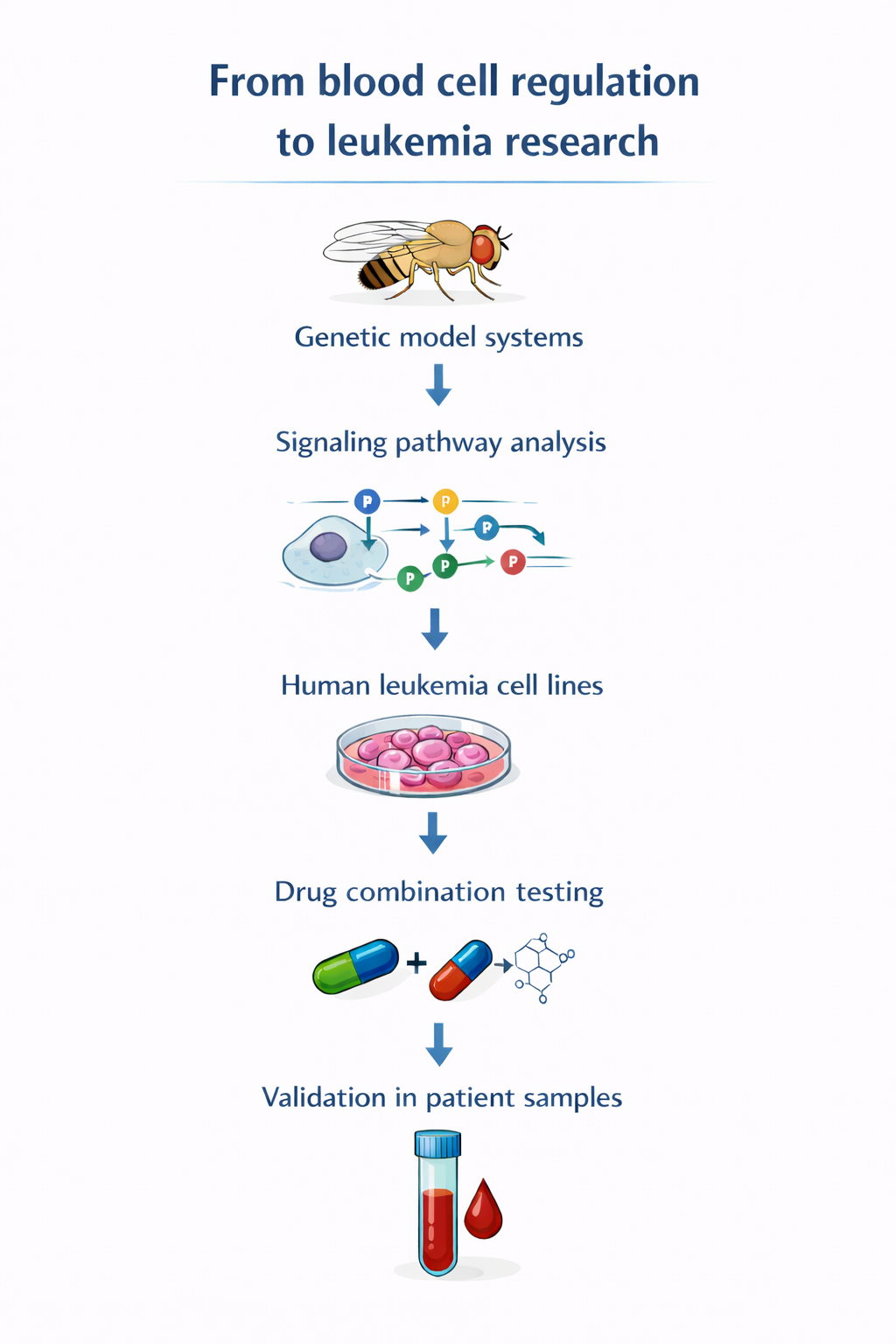
Our research seeks to better understand the behavior of leukemic and myeloproliferative cells and to identify molecular processes that could serve as therapeutic targets. We further investigate regulatory mechanisms discovered in genetic model systems using human leukemia cell lines, analyzing cancer cell growth and survival through cellular and molecular approaches. Additionally, we assess whether certain drugs are more effective in combination. The most promising drug candidates are then tested in patient blood samples, enabling evaluation in disease-relevant systems.
References:
Kharrat, B., Gábor, E., Vilmos, P., Goins, L. M., Honti, V. 2025. A novel role for Hsc70-4 in blood cell differentiation in Drosophila. Front Immunol. 16, 1641695. https://doi.org/10.3389/fimmu.2025.1641695
Kharrat, B., Gábor, E., Virág, N., Sinka, R., Jankovics, F., Kristó, I., Vilmos, P., Csordás, G., Honti V. 2024. Dual role for Headcase in hemocyte progenitor fate determination in Drosophila melanogaster. PLoS Genet. 20, e1011448. https://doi.org/10.1371/journal.pgen.1011448
Kúthy-Sutus, E., Kharrat, B., Gábor, E., Csordás, G., Sinka, R., Honti, V. 2022. A Novel Method for Primary Blood Cell Culturing and Selection in Drosophila melanogaster. Cells. 12, 24. https://doi.org/10.3390/cells12010024
Kharrat, B., Csordás, G., Honti, V. 2022. Peeling Back the Layers of Lymph Gland Structure and Regulation. Int J Mol Sci. 23, 7767. https://doi.org/10.3390/ijms23147767
Szkalisity, Á., Piccinini, F., Beleon, A., Balassa, T., Varga, I. G., Migh, E., Molnár, C., Paavolainen, L., Timonen, S., Banerjee, I., Ikonen, E., Yamauchi, Y., Andó, I., Peltonen, J., Pietiäinen, V., Honti, V., Horváth, P. 2021. Regression plane concept for analysing continuous cellular processes with machine learning. Nat Commun. 12, 2532. https://doi.org/10.1038/s41467-021-22866-x
Balog J. Á., Honti V., Kurucz, É., Kari, B., Puskás, L. G., Andó, I., Szebeni, G. J. 2021. Immunoprofiling of Drosophila Hemocytes by Single-cell Mass Cytometry. Genomics Proteomics Bioinformatics. 19, 243-252. https://doi.org/10.1016/j.gpb.2020.06.022
Csordás, G., Gábor, E., Honti, V., 2020. There and back again: The mechanisms of differentiation and transdifferentiation in Drosophila blood cells. Dev. Biol. 469, 135-143. https://doi.org/10.1016/j.ydbio.2020.10.006
Varga, G.I.B., Csordás, G., Cinege, G., Jankovics, F., Sinka, R., Kurucz, É., Andó, I., Honti, V., 2019. Headcase is a Repressor of Lamellocyte Fate in Drosophila melanogaster. Genes 10. https://doi.org/10.3390/genes10030173
Honti, V., Csordás, G., Kurucz, É., Márkus, R., Andó, I., 2014. The cell-mediated immunity of Drosophila melanogaster: hemocyte lineages, immune compartments, microanatomy and regulation. Dev. Comp. Immunol. 42, 47–56. https://doi.org/10.1016/j.dci.2013.06.005
Honti, V., Csordás, G., Márkus, R., Kurucz, E., Jankovics, F., Andó, I., 2010. Cell lineage tracing reveals the plasticity of the hemocyte lineages and of the hematopoietic compartments in Drosophila melanogaster. Mol. Immunol. 47, 1997–2004. https://doi.org/10.1016/j.molimm.2010.04.017
Honti, V., Kurucz, É., Csordás, G., Laurinyecz, B., Márkus, R., Andó, I., 2009. In vivo detection of lamellocytes in Drosophila melanogaster. Immunol. Lett. 126, 83–84. https://doi.org/10.1016/j.imlet.2009.08.004

senior research associate

research associate

research associate

PhD student

undergraduate student

undergraduate student

undergraduate student
 Viktor, HONTI
Viktor, HONTI
|
senior research associate | publications | CV |
 Erika, GÁBOR
Erika, GÁBOR
|
research associate | publications | CV |
 Zsanett, TAKÁCS
Zsanett, TAKÁCS
|
research associate | publications | CV |
 Dóra, BALOGH
Dóra, BALOGH
|
PhD student | publications | CV |
 Kata, CSOMOR
Kata, CSOMOR
|
undergraduate student | publications | CV |
 Kitti, PONGRÁCZ
Kitti, PONGRÁCZ
|
undergraduate student | publications | CV |
 Petra, VOZÁR
Petra, VOZÁR
|
undergraduate student | publications | CV |