
1. Myofibrils are composed of repeated sarcomeres that are extremely highly ordered macromolecular assemblies where structural organization is intimately linked to their functionality as contractile units. Recently, we developed a powerful nanoscopic approach that allowed us to determine the position of 27 muscle proteins with a quasi-molecular localization precision, and by means of template based protein structure modelling, we assembled a refined I-band and H-zone model with an unparalleled scope and resolution. We aim to combine this method with genetic approaches to explore the molecular mechanisms of sarcomere assembly during muscle development with special focus on the proteins involved in thin filament and Z-disc formation.
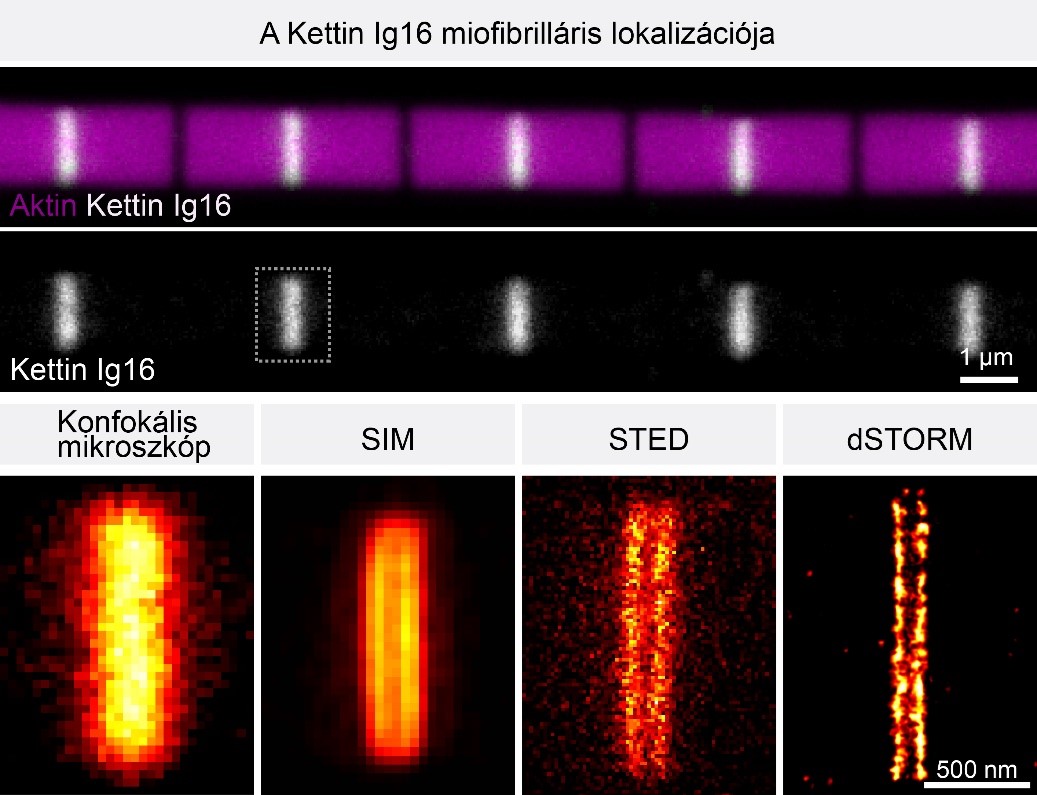
Comparison of the major superresolution microscopy methods reveals that the Z-disc protein Kettin appears as a single blurry band on confocal (CLSM) images, which is however, clearly resolved into two bands with SIM, STED and STORM
2. The highly dynamic actin cytoskeleton is one of the structurally and functionally most important cellular constituents. The actin cytoskeleton is involved in such fundamental cell biological processes as the maintenance of cell shape, cell division, intracellular transport, and motility. Cytoskeleton regulation involves the concerted action of a few hundred proteins. Of these, our research group mainly focuses on functional characterization of the formin type of proteins by studying their cellular and molecular mechanisms. Coordinated regulation of the actin and microtubule cytoskeleton is known to play a pivotal role in the growth and proper navigation of neuronal axons and dendrites that are necessary to the formation of a functional nervous system. One of our major scientific interests is to gain a better understanding of the cytoskeletal mechanisms of axonal growth and guidance by uncovering the role of formins that appear to play key roles in both actin and microtubule remodeling. Besides neuronal cytoskeleton regulation, we also investigate how formins contribute to cell shape changes and nuclear positioning during Drosophila compound eye development and oogenesis, respectively.
3. Within the framework of the National Laboratory for Biotechnology we aim to use our basic research results, gathered during our miofibrillogenesis studies, for the development of novel disease models. We expect that these efforts will help better understanding of certain muscle diseases and foster the development of novel therapeutic approaches.
Selected publications
Földi, I.; Tóth, K.; Gombos, R.; Gaszler, P.; Görög, P.; Zygouras, I.; Bugyi, B.; Mihály, J. Molecular Dissection of DAAM Function during Axon Growth in Drosophila Embryonic Neurons. CELLS 11 : 9 Paper: 1487 , 20 p. (2022)
Szikora, S.; Görög, P.; Mihály, J. The Mechanisms of Thin Filament Assembly and Length Regulation in Muscles. INTERNATIONAL JOURNAL OF MOLECULAR SCIENCES 23 : 10 Paper: 5306 (2022)
Chougule, A.; Lapraz, F.; Foldi, I.; Cerezo, D.; Mihaly, J.; Noselli, S. The Drosophila actin nucleator DAAM is essential for left-right asymmetry. PLOS GENETICS 16 : 4 Paper: e1008758 (2020)
Szikora, S., Gajdos, T., Novák, T., Farkas, D., Földi, I., Lenart, P., Erdélyi, M., Mihály, J. Nanoscopy reveals the layered organization of the sarcomeric H-zone and I-band complexes. J Cell Biol. 219(1): e201907026 (2020)
Migh E, Götz T, Földi I, Szikora S, Gombos R, Darula Z, Medzihradszky KF, Maléth J, Hegyi P, Sigrist S, Mihály J. Microtubule organization in presynaptic boutons relies on the formin DAAM. DEVELOPMENT 145 : 6 Paper: dev158519 , 13 p. (2018)
Villegas, SN.; Gombos, R.; Garcia-Lopez, L.; Gutierrez-Perez, I.; Garcia-Castillo, J.; Vallejo, DM.; Da, Ros VG.; Ballesta-Illan, E.; Mihaly, J.; Dominguez, M. PI3K/Akt Cooperates with Oncogenic Notch by Inducing Nitric Oxide-Dependent Inflammation. CELL REPORTS 22 : 10 pp. 2541-2549. , 9 p. (2018)
Szikora, S., Földi, I., Tóth, K., Migh, E., Vig, A., Bugyi, B., Maléth, J., Hegyi, P., Kaltenecker, P., Sanchez-Soriano, N., Mihály, J. The formin DAAM is required for coordination of the actin and microtubule cytoskeleton in axonal growth cones. J Cell Sci. 130(15):2506-2519. (2017)
Gombos, R ; Migh, E ; Antal, O ; Mukherjee, A ; Jenny, A ; Mihaly, J. The Formin DAAM Functions as Molecular Effector of the Planar Cell Polarity Pathway during Axonal Development in Drosophila. JOURNAL OF NEUROSCIENCE 35 : 28 pp. 10154-10167. , 14 p. (2015)
Molnár I, Migh E, Szikora S, Kalmár T, Végh AG, Deák F, Barkó S, Bugyi B, Orfanos Z, Kovács J, Juhász G, Váró G, Nyitrai M, Sparrow JC and Mihály J. DAAM is required for thin filament formation and sarcomerogenesis during muscle development in Drosophila. PLoS Genetics 10(2): e1004166 (2014)

scientific adviser

senior research associate

research associate

research associate

laboratory assistant

laboratory assistant
 József, MIHÁLY
József, MIHÁLY
|
scientific adviser | publications | CV |
 Szilárd, SZIKORA
Szilárd, SZIKORA
|
senior research associate | publications | CV |
 Balázs, VEDELEK
Balázs, VEDELEK
|
research associate | publications | CV |
 Dávid, FARKAS
Dávid, FARKAS
|
research associate | publications | CV |
 Anikó, BERENTE
Anikó, BERENTE
|
laboratory assistant | ||
 Ildikó, VELKEYNÉ KRAUSZ
Ildikó, VELKEYNÉ KRAUSZ
|
laboratory assistant |