
Cells are the building blocks of life. We use an interdisciplinary approach to explore the characteristics and functioning of various cell types, as well as their interactions with their microenvironment, and other cells. Furthermore, based on single cell level studies, we also aim to understand the collective behavior of cell populations and the functioning of complex cell communities.
We use advanced microfabrication, optical micromanipulation and microscopy techniques in order to trap single cells, observe them for extended periods of time and precisely engineer their microenvironment. We apply maskless photolithography based on laser direct writing, soft lithography and two-photon polymerization techniques to fabricate precisely engineered structures to trap cells and create their microenvironments [1] in the size range spanning from subcellular scales (250 nm) to several centimeters.
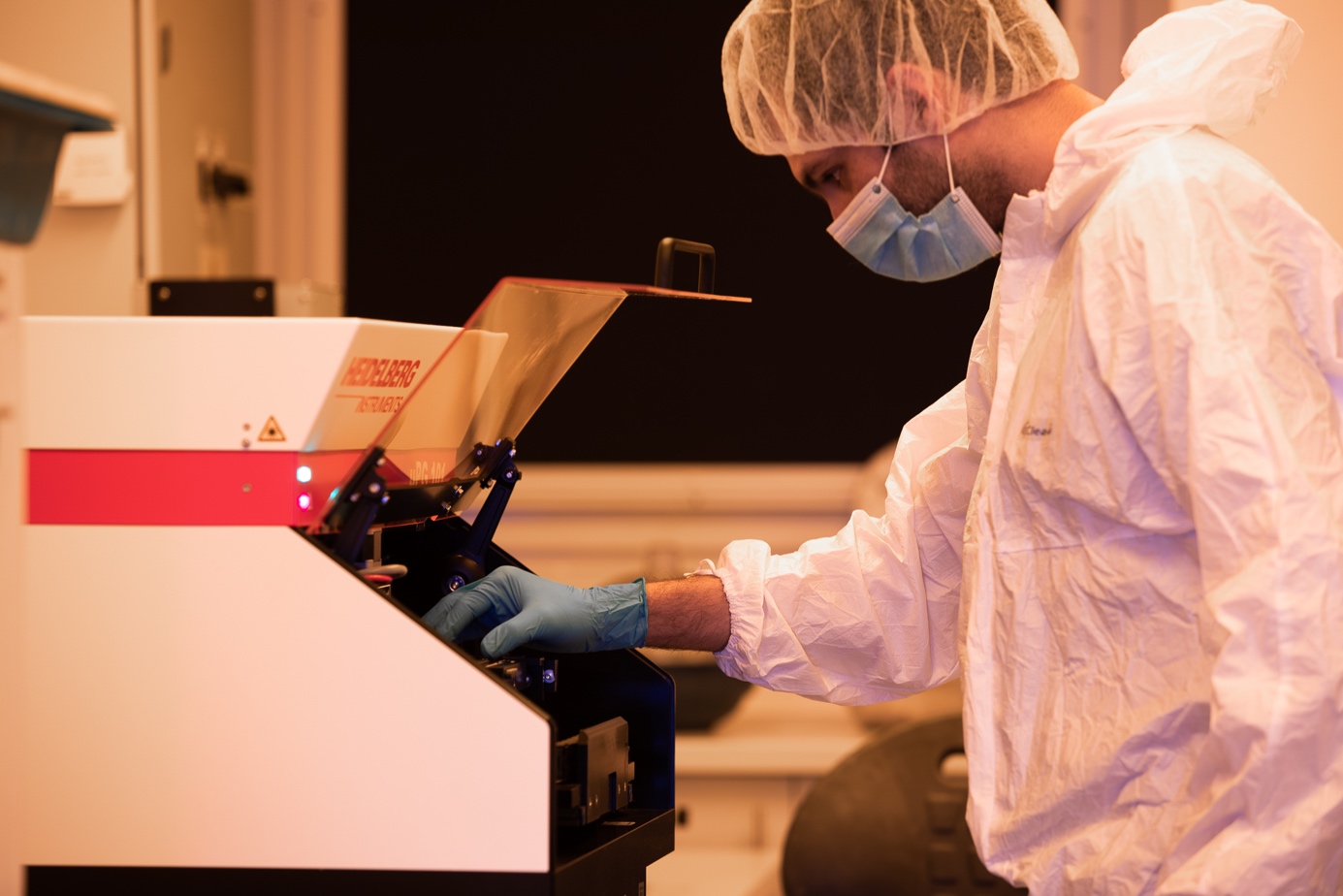

Microfabrication lab
We make microfluidic chips to trap single bacterial and algal cells, observe their properties and division, and explore the effect of cell microenvironment. With these chips we study the emergence of phenotypic heterogeneity (i.e. the variability of observable characteristics) in genetically identical cells. We develop microstructures that enable the trapping of cell progeny therefore we can analyze the correlation between relatedness and phenotype, as well as the effect of cellular aging.
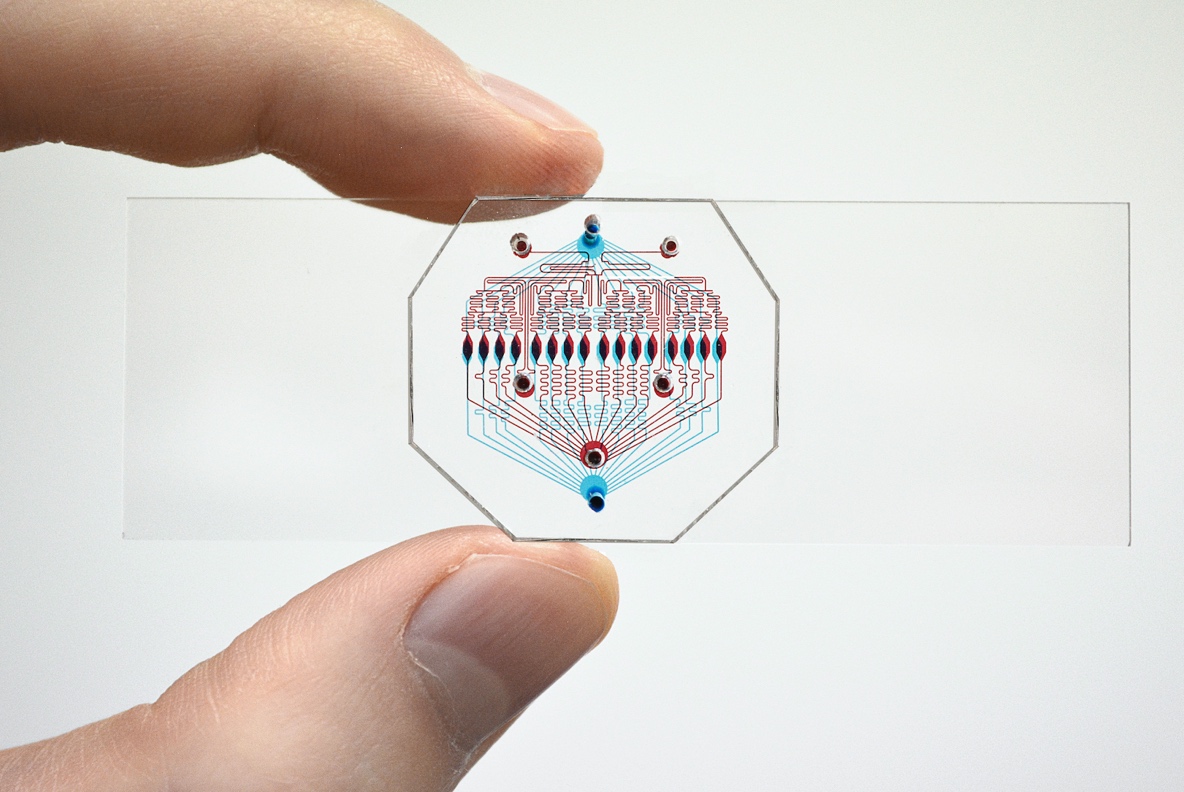

Microfluidic chip
Our group actively explore the possible applications of holographic optical trapping. We use dynamic holograms to create a trap system from an infrared laser beam to trap and precisely move microscopic particles [2] or even cells. With the combination of microfabricated devices bearing the size of a cell and the actuation with holographic optical tweezers, we are expanding the horizon of single cell manipulation and analysis. These microtools consist of two parts: a “handle” that interact with the trapping laser, and a “probe” interacting with the object of study.
In this line of studies, we used a microprobe as a mobile light guide [3] that we can aim to a particular spot on the cell to excite fluorescence locally. By moving a needle-shaped probe with the optical tweezers we can scan the surface of a cell, or we can explore the local mechanical properties. A needle probe covered by metal nanoparticles may be used for performing local tip-enhanced Raman spectroscopy [4] on the cell surface. We have recently developed a method to translate and rotate single cells in a fully controlled manner in order to observe them from different directions. Similarly to tomography, we can make 3D fluorescent images of cells that surpass conventional methods (e.g. confocal microscopy) in terms of axial resolution.
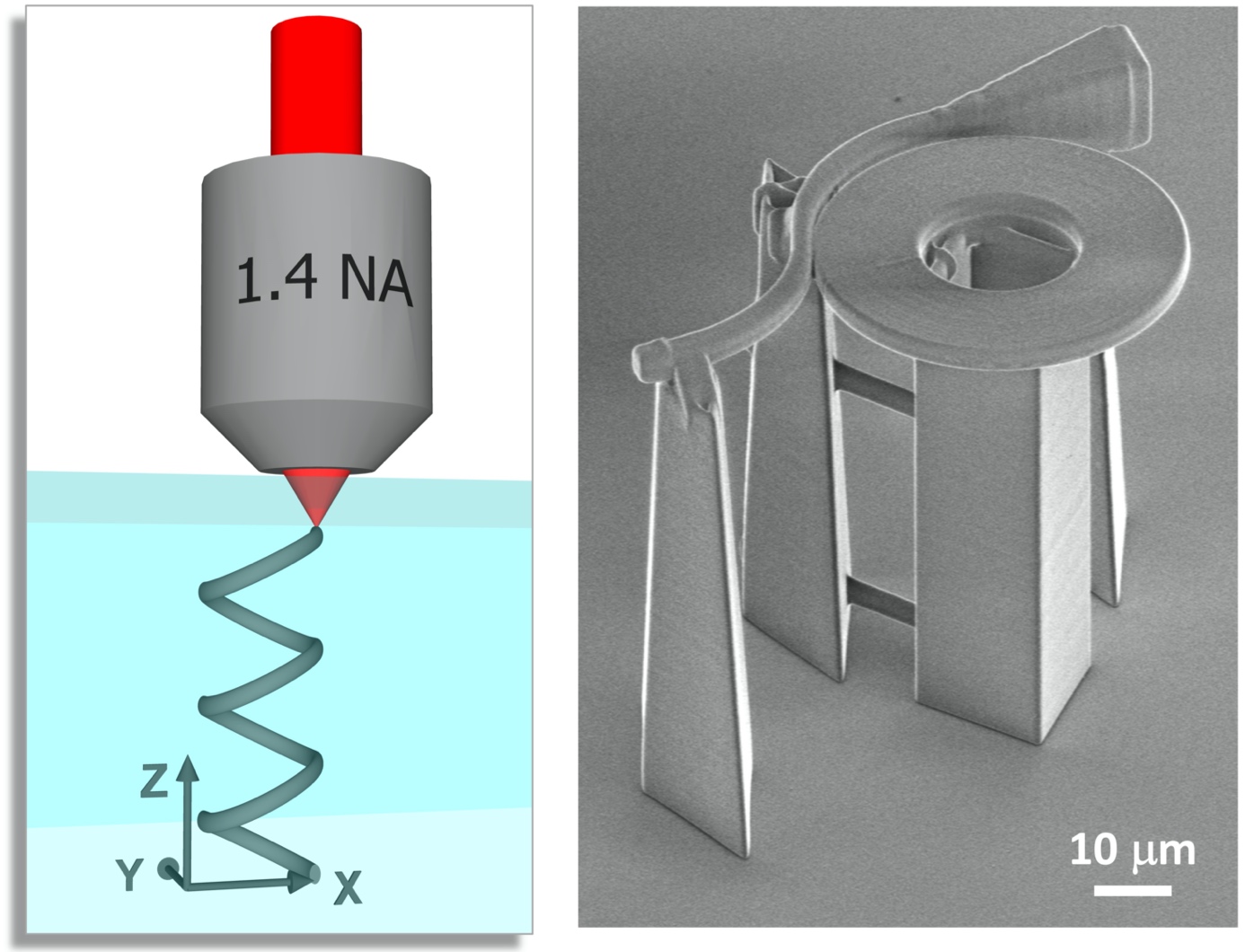
The schematics of photopolymerization and a scanning electronmicrograph of a fabricated protein detector structure.
We use microfabricated structures to study the hydrodynamic interactions of swimming cells with solid surfaces or other cells. For example we have demonstrated hydrodynamic trapping of swimming bacterial cells near concave surfaces [5], which promotes cell adhesion and biofilm formation. We used microfabricated propellers in holographic optical tweezers to explore hydrodynamic synchronization [6], which may be important for the correlated motion of bacterial flagellae and ciliae.
We use microfluidics to explore how individual properties and behavior of cells, as well as their interactions with each other and their environment lead to various phenomena on the population level [7]. We have developed a microfluidic platform to create stable spatial chemical gradients without liquid flow [8]. We use it to study bacterial chemotaxis, communication (quorum sensing) and evolution in heterogeneous environments. We apply a similar device to explore the interaction of physically separated but chemically coupled bacterial populations [9]. Our results help to understand the formation and functioning of multispecies microbial communities.
[1] Sima, F. et al. ACS Applied Bio Materials 1:1667-1676 (2018), DOI: 10.1021/acsabm.8b00487
[2] Vizsnyiczai, G. et al. SPIE PROC 9922:992216 (2016), DOI: 10.1117/12.2237601
[3] Palima, D. et al. Opt. Expr., 20: 2004-2014 (2012), DOI: 10.1364/OE.20.002004
[4] Vizsnyiczai, G. et al. Langmuir, 31:10087–10093 (2015), DOI: 10.1021/acs.langmuir.5b01210
[5] Nagy, K. et al. Front Microbiol 13 Paper: 820738, 12 p. (2022), https://doi.org/10.3389/fmicb.2022.820738
[6] Wetherington, M. ET al. BMC Biology 20:1 Paper: 262 (2022) https://doi.org/10.1186/s12915-022-01462-5
[7] Nagy, K. et al. Front Microbiol 9 p. 496 (2018) https://doi.org/10.3389/fmicb.2018.00496
[8] Nagy, K. Biomicrofluidics 9:4 Paper: 044105 (2015) https://doi.org/10.1063/1.4926981
[9] Fekete, T. et al. ACS Appl Mater Inter 13:33 p39018 (2021) https://doi.org/10.1021/acsami.1c08454
[10] Grexa, I. et al. Micromachines 11:9 Paper: 882 (2020) https://doi.org/10.3390/mi11090882
[11] Vizsnyiczai et al. Biomed Opt Express 11:2 p945 (2020) https://doi.org/10.1364/BOE.379233
[12] Kelemen, L. et al. LAB ON A CHIP 19:11 p1985 (2019) https://doi.org/10.1039/c9lc00174c
[13] Vizsnyiczai, G. Nat Commun 11:1 Paper: 2340 (2020) https://doi.org/10.1038/s41467-020-15711-0

senior research fellow, group leader

senior research fellow

research fellow

research fellow

research fellow

research fellow

scientific administrator

scientific administrator

PhD student

emeritus research professor
 Péter GALAJDA
Péter GALAJDA
|
senior research fellow, group leader | publications | CV |
 Lóránd KELEMEN
Lóránd KELEMEN
|
senior research fellow | publications | CV |
 Gaszton VIZSNYICZAI
Gaszton VIZSNYICZAI
|
research fellow | publications | CV |
 Krisztina NAGY
Krisztina NAGY
|
research fellow | publications | CV |
 András BUZÁS
András BUZÁS
|
research fellow | publications | CV |
 Tamás FEKETE
Tamás FEKETE
|
research fellow | publications | CV |
 Ágnes ÁBRAHÁM
Ágnes ÁBRAHÁM
|
scientific administrator | publications | CV |
 László DÉR
László DÉR
|
scientific administrator | publications | CV |
 Gergely T. IVÁNYI
Gergely T. IVÁNYI
|
PhD student | publications | CV |
 Pál ORMOS
Pál ORMOS
|
emeritus research professor | publications |